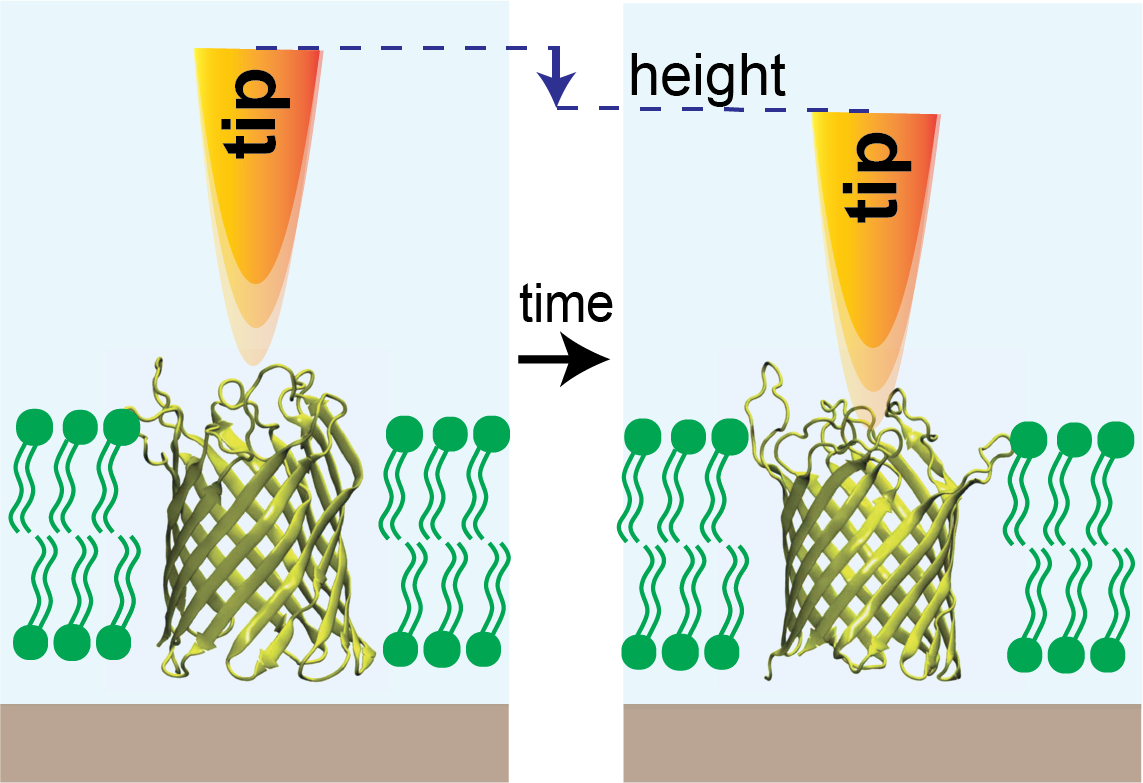
Membrane protein activity measured by high-speed atomic force microscopy (HS-AFM): A probe (oscillating tip shown in orange) hovering over the single closed ion channel in a membrane (left) descends into the channel (right) when it opens. This results in a detectable signal (change in height) over time similar to ion flow measured in single channel recordings. Image courtesy of Dr. Raghavendar Reddy Sanganna Gari.
Researchers at Weill Cornell Medicine have shown that they can record the high-speed motions of proteins while correlating their motion to function. The feat should allow scientists to study proteins in greater detail than ever before, and in principle enables the development of drugs that work better by hitting their protein targets much more effectively.
The researchers, in a study published July 16 in Nature Communications, adapted an imaging technique called high-speed atomic force microscopy (HS-AFM) and used it to record the rapid motions of a channel protein. Such proteins normally form channel- or tube-like structures in the membranes of cells, opening to allow the flow of molecules under certain conditions. The scientists were able to capture the rapid openings and closings of the channel protein at similar temporal resolution as single channel recordings, a standard technique to record the intermittent flow of charged molecules through the channel.
“There has been a strong need for a tool like this that reaches such a high bandwidth that it can ‘see’ the structural fluctuations of molecules as they work,” said senior author Dr. Simon Scheuring, professor of physiology and biophysics in anesthesiology at Weill Cornell Medicine.
Current techniques such as X-ray crystallography and electron microscopy allow researchers to make highly detailed images of molecules, revealing their structures down to atomic scale. These “pictures” typically are calculated from thousands of individual snapshots, and depict the average or dominant structural positionings, or conformations, of the molecules. They are, in that sense, like the long-exposure still images from the early days of photography.
However, many molecules are not rigid structures but, rather, flexible and always-moving machines. To reveal how such molecules move as they work, to see how their motion translates to function—to capture their key functional conformations, which might only exist fleetingly—scientists essentially need to make movies, not still images, and often very high-speed movies. Current techniques for dynamic structural imaging have many limitations, however, including, for many, the need to insert fluorescent tags on the molecules being imaged.
Dr. Scheuring and his laboratory have helped pioneer the use of HS-AFM, a tag-free method, to study molecular dynamics. The technique, which can image molecules in a liquid solution resembling a real cellular environment, features the use of a highly sensitive probe, much like the stylus of an old-fashioned record player, to feel its way across a molecule and thereby build up a picture of its structure. Standard HS-AFM isn’t fast enough to record the high-speed dynamics of many proteins, but Dr. Scheuring and colleagues have developed a modified version, HS-AFM height spectroscopy (HS-AFM-HS), that works much faster by capturing the dynamic changes of a target molecule in only one dimension, height.
In the new study led by first author Dr. Raghavendar Reddy Sanganna Gari, a postdoctoral research associate in Dr. Scheuring’s laboratory, the researchers used HS-AFM-HS to record the opening and closing of a relatively simple channel protein, OmpG, that is found in bacteria and is widely studied as a model channel protein. They were able to monitor the gating of OmpG at the effective rate of about 20,000 data points per second, watching how it shifted from open states to closed states or vice versa as they changed the acidity of the surrounding fluid.
More importantly, in collaboration with Dr. Crina Nimigean, professor of physiology and biophysics in anesthesiology, and her laboratory at WCM, they were able to correlate structural dynamics with functional dynamics for the first time in a membrane protein of this size.
The demonstration paves the way for the broader use of this technique in basic biology as well as pharmaceutical development.
“We’re now in an exciting period of HS-AFM technology, for example, using this technique to study how some drugs modulate the structural dynamics of the channel proteins they target,” said Dr. Sanganna Gari.
This story first appeared on the Weill Cornell Medicine Newsroom.

